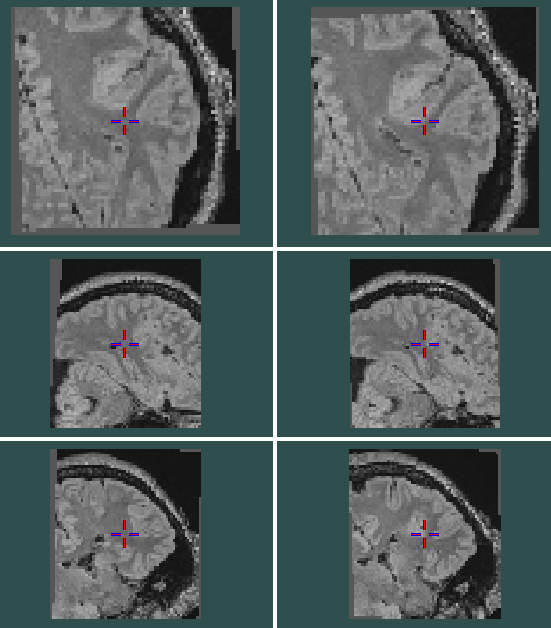
I’m comparing the result of SITK AffineTransform and Pytorch grid_sample. The difference between them is that sitk treats origin as the centre of rotation while Pytorch treats the centre of the image as the centre of rotation.
import SimpleITK as sitk
import numpy as np
import torch
import os
import pickle
import matplotlib.pyplot as plt
import copy
import imageio
import cv2
affine_matrix = np.array([[[ 1.0170, 0.0398, 0.0435, -0.0110],
[-0.0713, 1.0165, 0.0446, 0.0221],
[ 0.0387, 0.0189, 0.9905, -0.0033]]], dtype=np.float64)
torch_affine_matrix = torch.from_numpy(affine_matrix).unsqueeze(0)
img_3d = sitk.ReadImage("img.nii")
img_3d_tensor = torch.from_numpy(sitk.GetArrayFromImage(img_3d)).unsqueeze(0).unsqueeze(0)
# sitk transform
affine_3d_transform = sitk.AffineTransform(3)
affine_3d_transform.SetMatrix(theta_3d.squeeze().cpu().numpy()[:3, :3].flatten())
affine_3d_transform.SetTranslation(theta_3d.squeeze().cpu().numpy()[:, 3])
# affine_3d_transform.SetCenter((32, 32, 32)) # center of volume
# pytorch transform
resampled_3d_sitk = sitk.Resample(img_3d, img_3d, affine_3d_transform, sitk.sitkNearestNeighbor, 0.0)
sitk.WriteImage(resampled_3d_sitk, "sitk_3d_resampled.nii")
grid_3d = torch.nn.functional.affine_grid(theta_3d, img_3d_tensor.shape)
resampled_3d_pytorch = torch.nn.functional.grid_sample(img_3d_tensor, grid_3d)
resampled_3d_pytorch = sitk.GetImageFromArray(resampled_3d_pytorch.squeeze())
resampled_3d_pytorch.CopyInformation(img_3d)
sitk.WriteImage(resampled_3d_pytorch, "pytorch_3d_resampled.nii")
I’m not able to figure out why they are behaving differently. I might have missed something here. My assumption is that transforming the same volume by same matrix should give same result. Any help would be highly appreciated. Thanks in advance!
Result difference(left is SITK’s output and right is Pytorch’s output):
imge url: Dropbox - img.nii - Simplify your life