I need to create custom labels to draw shapes in a dicom.
It sounds like you want an interactive segmentation application. You should check out 3D Slicer and ITK-SNAP. Edit: link corrected.
If neither suit your needs, you need to explain your problem in much more detail.
I need to draw shapes in a dicom like you can do in itk-snap but I want to do it in python with simpleITK.
I need to know how to create a custom mask with simpleITK, for example to do something like this
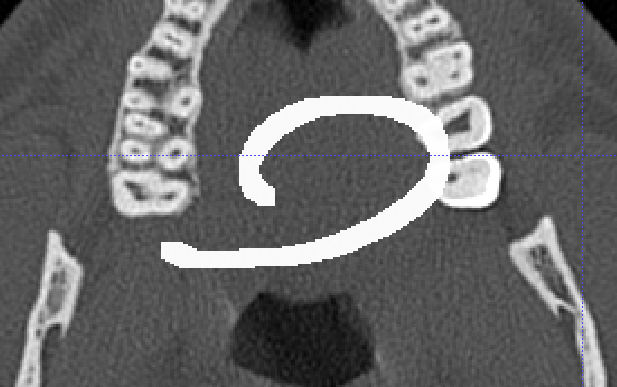
Hello @javi12,
There are two options when drawing:
- Modify the image itself.
- Overlay on the image (image serves as background and objects are overlaid onto it).
For the former case, please take a look at this Jupyter notebook. The approach of modifying the image itself only requires SimpleITK.
For the latter you can use matplotlib or any other plotting library with the image displayed in the background. This is what some of the GUI components that were developed for the notebooks do.
Hopefully one of these approaches solves your need.
Then 3D Slicer could be a good fit, as it provides a full Python3 environment: you can install any packages (SimpleITK is bundled by default), run your Python code, etc. and you an bring up a full-featured segmentation editor (with many manual, semi-automatic, and AI-assisted tools) when needed. Slicer also provides DICOM import/export, all kinds of visualization, processing, etc., so you don’t need to waste time with redeveloping basic things but you can spend time with implementing your own ideas. For example, if you are developing a segmentation tool that request some input from the user (seed points, painted regions, etc.) then you can implement it as Python scripted extension to 3D Slicer. If you prefer working with notebooks then you can also use Slicer as a Jupyter notebook kernel. If you have any questions about this then you can post it to 3D Slicer forum.