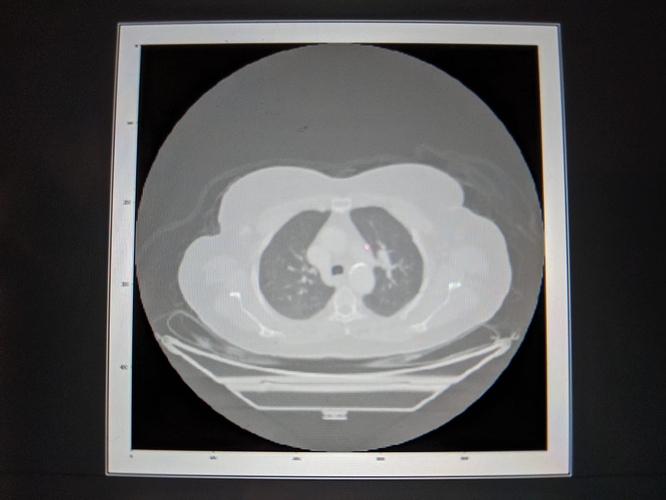
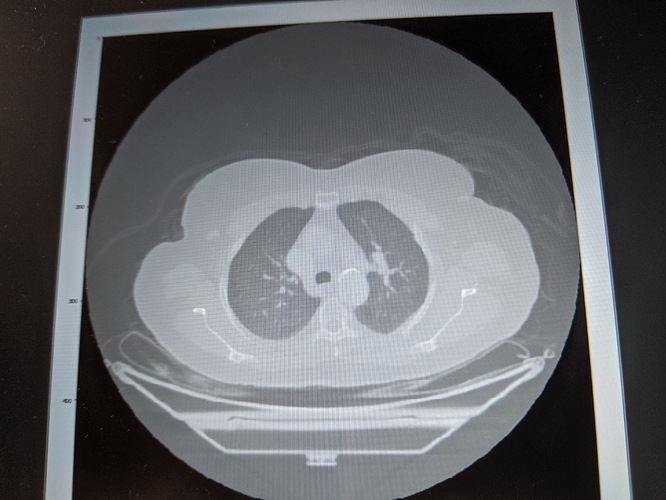
Hi All, I am working to detect my target in CT scan using simple ITK and python. I am forming list of seeds and passing it to confidence connected filter but what I observed is there are cases when as soon as I add one incorrect seed position in the list of seeds, confidence connected filter connects other targets surrounding my target of interest. Seems like this filter is too sensitive to list of seeds. Is there any other filter that provides connected component target across different 3D slices? Thanks,Jiten
Next filters in the region growing progression to study and understand would be “FastMarchingImageFilter”, and then “ThresholdSegmentationLevelSetImageFilter”.
Thanks @blowekamp This is good starting point I will try to understand and implement in my use case. Will share the observations and results.Thanks Jiten
Hi @blowekamp, I went over the concept of level set segmentation to understand Fast Marching filter, I have few questions.
Is the base 3D image itself considered as speed image in Fast Marching algorithm to control the front at the edges in 3D space?
Is it prerequisite to execute anisotropic diffusion and sigmoid filter before executing fast marching filter to get better results?
In the Level set segmentation technique, what does time actually signify, is the t value really represent time in seconds? I was not able to imagine intuition for time in level set segmentation. For example of I give time parameter value as 100 does it mean segmentation will run for 100 secs and whichever contours it was able to split and merge it will stop after 100 secs?
You can find a conventional way to run fast marching here:
https://simpleitk.readthedocs.io/en/master/Examples/FastMarchingSegmentation/Documentation.html
The 31_Levelset_Segmentation notebook also contains a widget to adjust the time threshold. You can visualized the contour or 0 value level set propagating over time. The speed input image gets integrated into the arrival time of the 0-value levelest which is initialized by the seeds. So, its not IRL time, but time of the differential equation that governs the system.
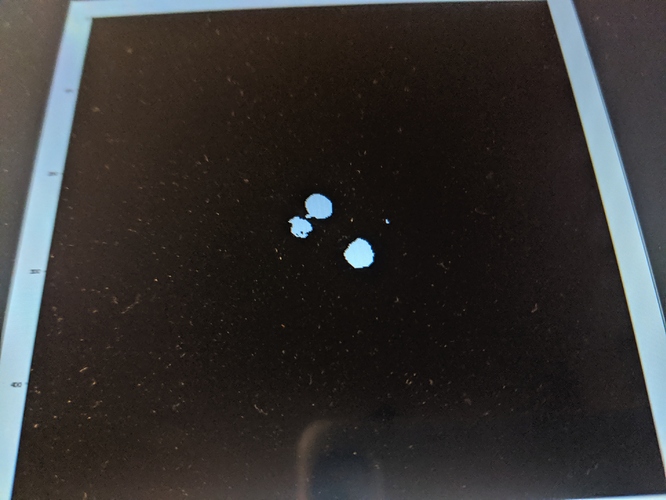
Hi @blowekamp I implemented the Fast Marching and results are pretty damn awesome!!! Much neater region growing technique. Attached is Chest CT segmented image using Fast Marching. If you look at the segmentation I am interested in segmenting only heart tissue within the Chest. This filter detected heart tissue between lungs very accurately. However I want to prevent the region surrounding heart region being segmented. Any ideas? I am further looking to understand parameters that I can tweak to just get heart tissue segmented. Thanks Jiten
Sample code snippet
ct_image_Smooth = sitk.CurvatureFlow(image1=ct_image, timeStep=0.125, numberOfIterations=5)
gradientMagnitude_output = sitk.GradientMagnitudeRecursiveGaussian(image1=ct_image_Smooth, sigma=1.0)
sigmoid_output = sitk.Sigmoid(image1=gradientMagnitude_output, alpha=-0.5, beta=3.0)
fastMarching = sitk.FastMarchingImageFilter()
trialPoint = (258, 250, 472)
fastMarching.AddTrialPoint(trialPoint)
trialPoint = (258, 280, 472)
fastMarching.AddTrialPoint(trialPoint)
trialPoint = (270, 277, 472)
fastMarching.AddTrialPoint(trialPoint)
trialPoint = (281, 216, 472)
fastMarching.AddTrialPoint(trialPoint)
trialPoint = (293, 247, 472)
fastMarching.AddTrialPoint(trialPoint)
trialPoint = (307, 233, 472)
fastMarching.AddTrialPoint(trialPoint)
fastMarching.SetStoppingValue(50)
fastMarchingOutput = fastMarching.Execute(sigmoid_output)
timeThreshold = 50
thresholder = sitk.BinaryThresholdImageFilter()
thresholder.SetLowerThreshold(0.0)
thresholder.SetUpperThreshold(timeThreshold)
thresholder.SetOutsideValue(0)
thresholder.SetInsideValue(255)
heartTissueLblImg = thresholder.Execute(fastMarchingOutput)
Hi @blowekamp Please find attached is the refined results I gained by setting the timethreshold and time stopping parameters to much lower value until it gave be very good segmentations of target of my interest! Thanks for the the guidance. Level Set technique of region growing is much more reliable and robust. Any ideas on which edge identification filters goes hand in hand with Fast marching level set filter? I observed that if we apply some good edge detection filter to make contour edges sharper, the results of fast marching are much much enhanced!! Currently I am using Curvature flow for smoothing prior to applying gradient magnitude recursive gaussian, sigmoid and finally fast marching in sequence. Thanks,Jiten