Hello,
I want to register two 3D multi-modal medical images using mutual information. However instead of using the inbuilt MattesMutualInformation or JointHistogramMutualInformation, I want to use my own mutual information computation function. The rest of the code, as given in SITK, will remain unchanged. Is this possible? If yes, how can I do that?
To define your own metric, you might have to implement it in ITK and expose it in SimpleITK. @machadoL wanted to write his own variant of mutual information, he might update us on progress and maybe share his experience.
Thank you @dzenanz !! Following your advice and the chain of conversation you mentioned, I have found that the computation of mutual information is happening here. Please correct me if I am wrong.
Now I want to run this code, preferably in debugger mode, and modify it. How can I do that? My knowledge in C++ is limited to high school level.
The easiest way is to configure ITK to build with examples (BUILD_EXAMPLES=ON), compile it in debug mode, and run one of the registration examples which uses mutual information (such as ImageRegistration13). Put a breakpoint somewhere in the code you highlighted, and see what happens. If you modify that code, most IDEs (integrated development environments) should automatically recompile the example you are using.
Thank you @dzenanz . I have configured ITK.
I have run Example codes ImageRegistration1.cxx to ImageRegistration20.cxx putting breakpoints in itkJointHistogramMutualInformationImageToImageMetricv4.hxx. However, none of the codes stopped at the breakpoints.
Please tell me which code calls itkJointHistogramMutualInformationImageToImageMetricv4.hxx.
Also, I could not run the codes which require 3D images as argument (ImageRegistration8.cxx and ImageRegistration20.cxx). The following error is showing.
Could not create IO object for reading file: C:\Users\...BrainWeb\brainweb1e1a10f20.mha. Tried to create one of the following: BMPImageIO BioRadImageIO Bruk..
How can I resolve this?
Not all examples use mutual information. Example1 uses mean squares, and so does example 20. Make sure that the example you try uses mutual information, and not another metric type.
BMPImageIO BioRadImageIO Bruk…
What is the complete list of ImageIOs tried? Is MetaIO or MetaImageIO on that list? That’s what .mha file should be read by.
msg | 0x0000021c9d765d40 Could not create IO object for reading file C:\\Users\\ …\\ITK1\\Examples\\Data\\BrainWeb1\\brainweb1e1a10f20.mha\n Tried to create one of the following:\n BMPImageIO\n BioRadImageIO\n Bruk... | std::basic_ostringstream<char,std::char_traits<char>,std::allocator<char>>
This is the entire error message. How can I find out the complete list of ImageIOs? I am using Visual Studio 2019.
I tried to set the ImageIO with the following command, but it did not work.
fixedImageReader->SetImageIO("MetaImageIO");
Apparently, SetImageIO expects itk::ImageIOBase* as the datatype of the input parameter. I do not know how to convert "MetaImageIO" to itk::ImageIOBase*.
itk::MetaImageIO::Pointer metaIO = itk::MetaImageIO::New();
fixedImageReader->SetImageIO(metaIO);
Thank you @zivy . I am able to read 3D images with these commands.
I have modified itkJointHistogramMutualInformationImagetoImageMetricv4.cxx. The code is now computing mutual information according to the modified formula. I have used the modified formula to register two 3D images (CT and MR), and it is giving acceptable result.
Now I want to expose this modification to SITK. Please tell me the process.
Hello @debapriya,
To expose the new ITK metric in SimpleITK, add to the existing metrics by following the existing pattern. The relevant locations: CreateMetric and the SetMetricAsX.
Thank you @zivy .
I saved my modified code as itkJointHistogramMutualInformationImageToImageEfficientEntropyMetricv4.hxx.
I configured and generated ITK using CMake and built solution in Visual Studio 2019.
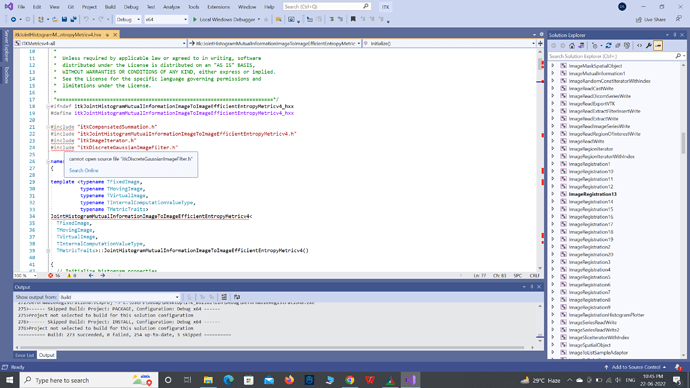
Now, when I open itkJointHistogramMutualInformationImageToImageEfficientEntropyMetricv4.hxx, it says it cannot open any source file except itkJointHistogramMutualInformationImageToImageEfficientEntropyMetricv4.h.
Apparently, it cannot find the source files which are located in other folders.
What step am I missing?
Below is a screenshot of the message
Hello @debapriya,
I suspect this is an issue with the VS IDE configuration files generated via CMake but really don’t know. As this is on windows, possibly close the program and open it again?
Possibly @dzenanz or @matt.mccormick may have an idea.
I deleted the ITK_build folder and re-configured ITK afresh. However, the same problem persists. Not only my newly added file itkJointHistogramMutualInformationImageToImageEfficientEntropyMetricv4, but also existing files like itkJointHistogramMutualInformationImageToImageMetricv4 and itkImageRegistrationMethodv4 are showing the same error.
I tried to run one of the Example codes with breakpoint, but the code is bypassing the lines where itkImageRegistrationMethodv4 or itkJointHistogramMutualInformationImageToImageMetricv4 are called.
These codes were running perfectly well before I configured itkJointHistogramMutualInformationImageToImageEfficientEntropyMetricv4.
Please tell me what is happening and how to resolve this.
Please tell me the process of adding new .h and .hxx files in the ITK\Modules\Registration\Metricsv4\include folder and compiling them. What all files need to be updated?
using MetricType =
itk::JointHistogramMutualInformationImageToImageEfficientEntropyMetricv4<
FixedImageType,
MovingImageType>;
When I add the above code snippet, I get a message namespace *itk* has no member *JointHistogramMutualInformationImageToImageEfficientEntropyMetricv4*.
How can I add JointHistogramMutualInformationImageToImageEfficientEntropyMetricv4 to namespace itk?
Or, is there a process to add and compile .h or .hxx files from outside and call them in the codes?
Hello @debapriya ,
Because this is a templated class the code isn’t compiled when you build ITK, so you don’t really know if the syntax is even valid. What you need to do when adding a new metric to ITK:
- Make sure that the class is indeed in the itk namespace (like this), and in the expected directory.
- Add a test which instantiates the template. The compiler will create the code and validate that the syntax is correct (like this).
- Add the test to the CMakeLists.txt file (see here).
- Build ITK with testing turned on.
Thank you so much @zivy . Finally, I could make some progress with your help. However, I am getting errors while building ITK following your steps. My lack of knowledge in advanced C++ is making things worse. Probably, I’ll go through some advanced C++ tutorial and try this again.
Can you clear a basic doubt for me?
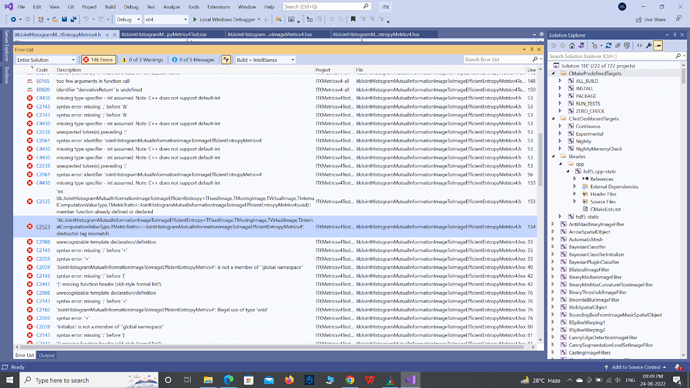
My metric itkJointHistogramMutualInformationImageToImageEfficientEntropyMetricv4.hxx and itkJointHistogramMutualInformationImageToImageEfficientEntropyMetricv4.h are photocopies of itkJointHistogramMutualInformationImageToImageMetricv4.hxx and itkJointHistogramMutualInformationImageToImageMetricv4.h, with modifications done here and here respectively. The modification consists of algebraic computations only. How come itkJointHistogramMutualInformationImageToImageMetricv4.hxx is compiled errorless while itkJointHistogramMutualInformationImageToImageEfficientEntropyMetricv4.hxx is throwing errors in huge numbers?
Below is a screenshot of the list of errors.
Hello @zivy,
Finally some good news!! My code compiled without error!! All those huge list of errors in the above screenshot was due to some silly mess that I had created. Didn’t take advanced C++ knowledge to debug ![]() . Big thanks to you.
. Big thanks to you.
Now I have to expose this code in SimpleITK. New challenge ahead ![]() !!
!!
Hello,
I am trying to expose my new ITK metric in SimpleITK. However, when I add
#include "itkMutualInformationImageToImageEfficientEntropyMetricv4.h" to sitkImageRegistrationMethod_CreateMetric.hxx, I get the following two errors.
-
Cannot open include file: 'itkMutualInformationImageToImageEfficientEntropyMetricv4.h': No such file or directory [C:\Users\...\Desktop\SITK_build\SimpleITK-build\Code\Registration\src\SimpleITKRegistration.vcxproj] -
Custom build for 'C:\Users\...\Desktop\SITK_build\CMakeFiles\6ac7ef3cb40dea1d1e5f86faaa9d2a21\SimpleITK-configure.rule;C:\Users\...\Desktop\SITK_build\CMakeFiles\6ac7ef3cb40dea1d1e5f86faaa9d2a21\SimpleITK-forcebuild.rule;C:\Users\...\Desktop\SITK_build\CMakeFiles\6ac7ef3cb40dea1d1e5f86faaa9d2a21\SimpleITK-build.rule;C:\Users\...\Desktop\SITK_build\CMakeFiles\6ac7ef3cb40dea1d1e5f86faaa9d2a21\SimpleITK-install.rule;C:\Users\...\Desktop\SITK_build\CMakeFiles\afb7cf371e783ad4daf13950766043ce\SimpleITK-complete.rule;C:\Users\...\Desktop\SITK_build\CMakeFiles\ca4cabce27f87d6108ded18de02ff787\SimpleITK.rule' exited with code 1.
Earlier, I have compiled itkMutualInformationImageToImageEfficientEntropyMetricv4.h in ITK successfully, following the steps given by @zivy and have used it to debug and run Example image registration codes in ITK.
Please help me resolve this.
P.S. I have built SimpleITK on the SuperBuild sub-directory. Could that be a reason for the error?
Hello @debapriya,
Please follow the SimpleITK build instructions on read-the-docs. Take the Building Manually path, as you need to point to your modified ITK version.
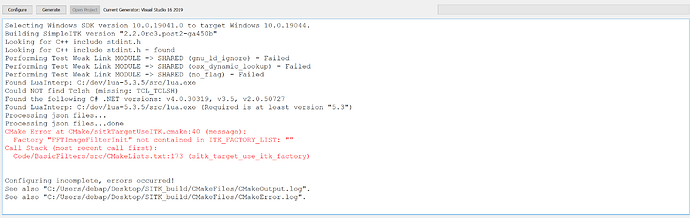
I am trying to build SimpleITK using the Building Manually path. I am getting the following error in CMake
Looks like ITK_FACTORY_LIST is empty, but I do not know how to populate it. Please help.