Hi!
I’ve recently started using the Python bindings for the latest version of ITK and it is working really great so far!
I’m currently building an interactive application for testing effects of different filters with my own reconstructed MRI data. However, when I’m using the itkwidgets view function to look at the data I’m noticing some strange behaviour when I’m updating the current image, the apparent intensity levels in the image changes, eventhough the image haven’t changed.
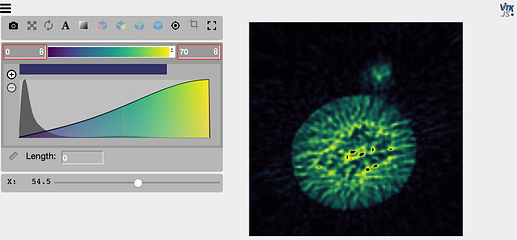
Example: I have produced an image as an itk image object and view it as
viewer = view(img, mode='x', vmin=0, vmax=70)
viewer
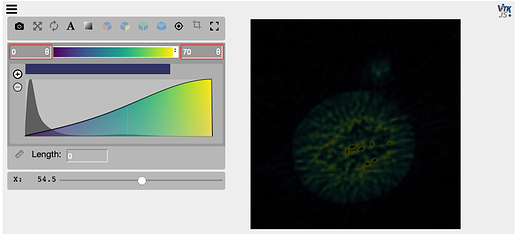
Now I add in another line to just update the viewer with the same image again, as I need to do when testing out different filters interactively, and the image comes out much darker, eventhough I have the same window levels. Any ideas why?
viewer = view(img, mode='x', vmin=0, vmax=70)
viewer.image = img # <-- New line
viewer
For debug purpose
type(img)
>>> itk.itkImagePython.itkImageD3
What I didn’t show here is that the viewer will also override the values I set for vmin and vmax when I update the image.
I got the syntax for updating the current image (viewer.image = image) from your tutorial 3 on filtering at https://github.com/KitwareMedical/CourseInBiomedicalImageAnalysisVisualizationAndArtificialIntelligence
Is there something I’m missing in terms of updating the current image?
Thank you very much!