Greetings!
I am a uni student relatively new to image/file processing. I am encountering something which I view as an issue during the processing of my 3D images. For the record, what I do with the original DICOM images, is that I turn them into numpy arrays, and combine arrays corresponding to different slice locations into a volume. I have a number of these 3D arrays, which thus form a time dependent dataset. These volumes are stored in a folder in an input direcotry, and I would like to register them using SimpleITK’s functions. This is my code:
import os
import numpy as np
import SimpleITK as sitk
import shutil
def register_images(input_dir, output_dir):
# Create the output directory if it doesn't exist
if not os.path.exists(output_dir):
os.makedirs(output_dir)
# Get a list of all files in the input directory
file_list = os.listdir(input_dir)
file_number_str = [x[7:-4] for x in file_list]
file_number_int = list(map(int, file_number_str))
file_number_int_sorted, file_list = (list(t) for t in zip(*sorted(zip(file_number_int, file_list))))
volumes = []
# Load the first image as the reference image
ref_array = np.load(os.path.join(input_dir, file_list[0]))
ref_image = sitk.GetImageFromArray(ref_array)
# Copy the reference volume to the output directory
ref_output_path = os.path.join(output_dir, file_list[0])
shutil.copy(os.path.join(input_dir, file_list[0]), ref_output_path)
print(f"Reference volume copied to: {ref_output_path}")
# Set up the registration parameters
registration_method = sitk.ImageRegistrationMethod()
registration_method.SetMetricAsMeanSquares()
registration_method.SetOptimizerAsRegularStepGradientDescent(learningRate=0.1, minStep=1e-4, numberOfIterations=1000)
registration_method.SetInitialTransform(sitk.Euler3DTransform())
# Register each subsequent image to the reference image
for file_name in file_list[1:]:
file_path = os.path.join(input_dir, file_name)
moving_array = np.load(file_path)
moving_image = sitk.GetImageFromArray(moving_array)
# Perform registration
final_transform = registration_method.Execute(ref_image, moving_image)
# Apply the transform to the moving image
registered_image = sitk.Resample(moving_image, ref_image, final_transform, sitk.sitkLinear, 0, moving_image.GetPixelID())
# Save the registered image to the output directory
output_path = os.path.join(output_dir, file_name)
registered_array = sitk.GetArrayFromImage(registered_image)
np.save(output_path, registered_array)
print(f"Registered image saved: {output_path}")
print("Registration and saving completed successfully.")
input_directory = r'path\to\input\folder'
output_directory = r'path\to\output\folder'
register_images(input_directory, output_directory)
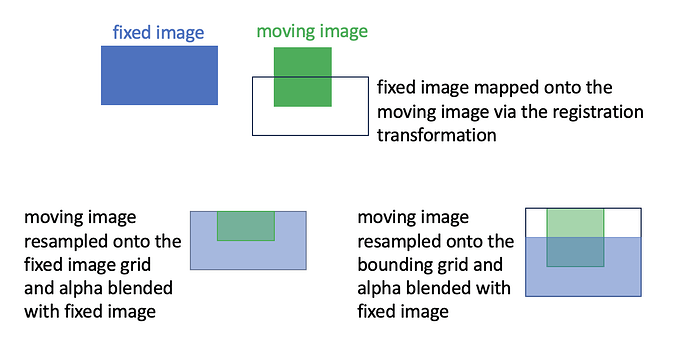
The problem is that parts of the images seem to vanish after I try yo plot them (or make an animation to showcase the registration). Is this something that is a result of the resampling, the registration method, or is there fundamentally something wrong with my code?
Thanks in advance for your response, have a great day!
PS: this is what some of the images that miss parts look like, so that you can you see what I mean:
Good image
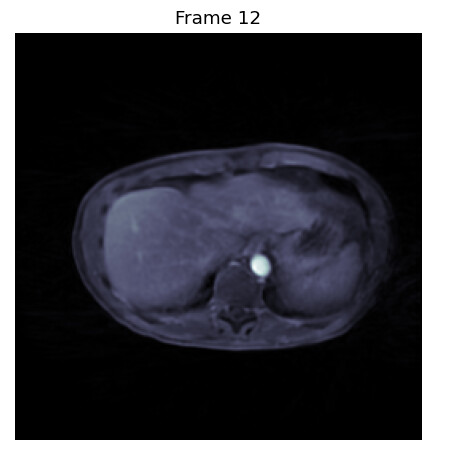
Bad images