Hi mat,
I am trying to build and run WatershedSegmentation1.cxx using ITK-WASM/WASI.
the code is below. I have just tried to parse input command line with pipeline as in example InputsOutputs example, but with one exception that i am trying to parse file names for input image file name and output image file name instead of inputImage and outputImage. It does build without any errors. when i try to run the WatershedSegmentation.wasi.wasm as follows:
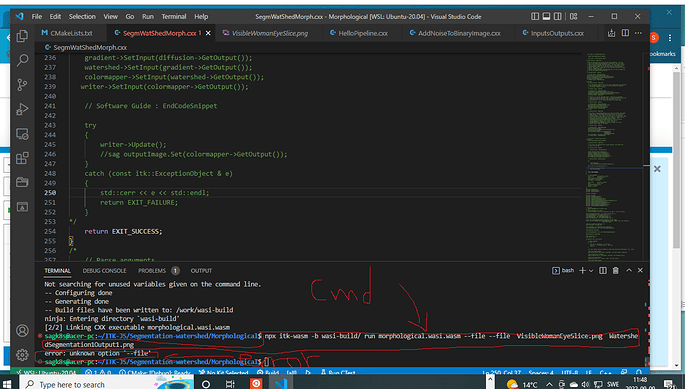
npx itk-wasm -b wasi-build run WatershedSegmentation1.wasi.wasm --file --file VisibleWomeEyeSlice.png WatershedSegementation1Output1.png
it gives me error that --file unknown option?!
can you help me to solve this issue. I have commented out rest of the code as you can see in the code listing above. I firstly try to parse the file and print them from pipeline in command line. not working,
I wrote the code another way by passing the images of input image and outputimage just like InputsOutputs example provided by you in ITK-WASM page. I could parse the images but i cannot get to write the segmented image using the outputImage. outputImage.Set(colormapper->GetOutput());did parse input image and output image but it crahses onSet(colormapper->GetOutput());`
`
// Software Guide : BeginCommandLineArgs
// INPUTS: {VisibleWomanEyeSlice.png}
// OUTPUTS: {WatershedSegmentation1Output1.png}
// ARGUMENTS: 2 10 0 0.05 1
// Software Guide : EndCommandLineArgs
// Software Guide : BeginCommandLineArgs
// INPUTS: {VisibleWomanEyeSlice.png}
// OUTPUTS: {WatershedSegmentation1Output2.png}
// ARGUMENTS: 2 10 0.001 0.15 0
// Software Guide : EndCommandLineArgs
// Software Guide : BeginLatex
//
// The following example illustrates how to preprocess and segment images
// using the \doxygen{WatershedImageFilter}. Note that the care with which
// the data are preprocessed will greatly affect the quality of your result.
// Typically, the best results are obtained by preprocessing the original
// image with an edge-preserving diffusion filter, such as one of the
// anisotropic diffusion filters, or the bilateral image filter. As
// noted in Section~\ref{sec:AboutWatersheds}, the height function used as
// input should be created such that higher positive values correspond to
// object boundaries. A suitable height function for many applications can
// be generated as the gradient magnitude of the image to be segmented.
//
// The \doxygen{VectorGradientMagnitudeAnisotropicDiffusionImageFilter} class
// is used to smooth the image and the
// \doxygen{VectorGradientMagnitudeImageFilter} is used to generate the
// height function. We begin by including all preprocessing filter header
// files and the header file for the WatershedImageFilter. We
// use the vector versions of these filters because the input dataset is a
// color image.
//
//
// Software Guide : EndLatex
#include “itkPipeline.h”
#include “itkInputImage.h”
#include “itkOutputImage.h”
#include “itkImage.h”
#include <iostream>
// Software Guide : BeginCodeSnippet
#include “itkVectorGradientAnisotropicDiffusionImageFilter.h”
#include “itkVectorGradientMagnitudeImageFilter.h”
#include “itkWatershedImageFilter.h”
// Software Guide : EndCodeSnippet
#include “itkImageFileReader.h”
#include “itkImageFileWriter.h”
#include “itkCastImageFilter.h”
#include “itkScalarToRGBPixelFunctor.h”
int main(int argc, char* argv[])
{
constexpr unsigned int Dimension = 2;
constexpr unsigned int VDimension = 3;
// Software Guide : BeginCodeSnippet
using RGBPixelType = itk::RGBPixel;
using RGBImageType = itk::Image<RGBPixelType, Dimension>;
using VectorPixelType = itk::Vector<float, VDimension>;
using VectorImageType = itk::Image<VectorPixelType, Dimension>;
using LabeledImageType = itk::Image<itk::IdentifierType, Dimension>;
using ScalarImageType = itk::Image<float, Dimension>;
using InputImageType = itk::wasm::InputImage;
using OutputImageType = itk::wasm::OutputImage;
//initialization of variables
unsigned int conductanceTerm = 2;
unsigned int diffusionIterations = 10;
double lowerThreshold = 0.0;
double outputScaleLevel = 0.05;
unsigned int gradientMode = 1;
std::string inputFileName = "";
std::string outputFileName = "";
itk::wasm::Pipeline pipeline("Segment input image using Watershed method itk::wasm::pipeline", argc, argv);
//Add input image argument
InputImageType inputImage;
pipeline.add_option("-f, --file", inputFileName, "the input image")->required();
//Add input image argument
OutputImageType outputImage;
pipeline.add_option("-f, --file", outputFileName, "the output image")->required();
//Add conductanceTerm value argument
pipeline.add_option("-c, --conductanceTerm", conductanceTerm, "the conductanceTerm value");
//Add diffusion iterations value
pipeline.add_option("-d, --diffusionIterations", diffusionIterations, "the diffusionIterations value");
pipeline.add_option("-l, --lowerThreshold", lowerThreshold, "the lowerThreshold value");
//Add outputScaleLevel value
pipeline.add_option("-o, --outputScaleLevel", outputScaleLevel, "the outputScaleLevel value");
//Add gradientMode value
pipeline.add_option("-g, --gradientMode", gradientMode, "the gradientMode value");
//parse the pipeline input
ITK_WASM_PARSE(pipeline);
std::cout << "input File Name: " << inputFileName << std::endl;
std::cout << "output File Name: " << outputFileName << std::endl;
/*
// Software Guide : BeginCodeSnippet
using FileReaderType = itk::ImageFileReader;
using CastFilterType = itk::CastImageFilter<RGBImageType, VectorImageType>;
using DiffusionFilterType =
itk::VectorGradientAnisotropicDiffusionImageFilter<VectorImageType,
VectorImageType>;
using GradientMagnitudeFilterType =
itk::VectorGradientMagnitudeImageFilter;
using WatershedFilterType = itk::WatershedImageFilter;
// Software Guide : EndCodeSnippet
using FileWriterType = itk::ImageFileWriter<RGBImageType>;
auto reader = FileReaderType::New();
reader->SetFileName(inputFileName);
auto caster = CastFilterType::New();
// Software Guide : BeginLatex
//
// Next we instantiate the filters and set their parameters. The first
// step in the image processing pipeline is diffusion of the color input
// image using an anisotropic diffusion filter. For this class of filters,
// the CFL condition requires that the time step be no more than 0.25 for
// two-dimensional images, and no more than 0.125 for three-dimensional
// images. The number of iterations and the conductance term will be taken
// from the command line. See
// Section~\ref{sec:EdgePreservingSmoothingFilters} for more information on
// the ITK anisotropic diffusion filters.
//
// Software Guide : EndLatex
// Software Guide : BeginCodeSnippet
auto diffusion = DiffusionFilterType::New();
diffusion->SetNumberOfIterations(diffusionIterations);
diffusion->SetConductanceParameter(conductanceTerm);
diffusion->SetTimeStep(0.125);
// Software Guide : EndCodeSnippet
//sag check
// Software Guide : BeginLatex
//
// The ITK gradient magnitude filter for vector-valued images can optionally
// take several parameters. Here we allow only enabling or disabling
// of principal component analysis.
//
// Software Guide : EndLatex
// Software Guide : BeginCodeSnippet
auto gradient = GradientMagnitudeFilterType::New();
//gradient->SetUsePrincipleComponents(std::stoi(gradientMode));
gradient->SetUsePrincipleComponents(gradientMode);
// Software Guide : BeginLatex
//
// Finally we set up the watershed filter. There are two parameters.
// \code{Level} controls watershed depth, and \code{Threshold} controls the
// lower thresholding of the input. Both parameters are set as a
// percentage (0.0 - 1.0) of the maximum depth in the input image.
//
// Software Guide : EndLatex
// Software Guide : BeginCodeSnippet
auto watershed = WatershedFilterType::New();
watershed->SetLevel(outputScaleLevel);
watershed->SetThreshold(lowerThreshold);
// Software Guide : EndCodeSnippet
//sag check
// Software Guide : BeginLatex
//
// The output of WatershedImageFilter is an image of unsigned long integer
// labels, where a label denotes membership of a pixel in a particular
// segmented region. This format is not practical for visualization, so
// for the purposes of this example, we will convert it to RGB pixels. RGB
// images have the advantage that they can be saved as a simple png file
// and viewed using any standard image viewer software. The
// \subdoxygen{Functor}{ScalarToRGBPixelFunctor} class is a special
// function object designed to hash a scalar value into an
// \doxygen{RGBPixel}. Plugging this functor into the
// \doxygen{UnaryFunctorImageFilter} creates an image filter which
// converts scalar images to RGB images.
//
// Software Guide : EndLatex
// Software Guide : BeginCodeSnippet
using ColormapFunctorType =
itk::Functor::ScalarToRGBPixelFunctor<unsigned long>;
using ColormapFilterType =
itk::UnaryFunctorImageFilter<LabeledImageType,
RGBImageType,
ColormapFunctorType>;
auto colormapper = ColormapFilterType::New();
// Software Guide : EndCodeSnippet
auto writer = FileWriterType::New();
//sag check
writer->SetFileName(outputFileName);
// Software Guide : BeginLatex
//
// The filters are connected into a single pipeline, with readers and
// writers at each end.
//
// Software Guide : EndLatex
// Software Guide : BeginCodeSnippet
caster->SetInput(reader->GetOutput());
diffusion->SetInput(caster->GetOutput());
gradient->SetInput(diffusion->GetOutput());
watershed->SetInput(gradient->GetOutput());
colormapper->SetInput(watershed->GetOutput());
writer->SetInput(colormapper->GetOutput());
// Software Guide : EndCodeSnippet
try
{
writer->Update();
//sag outputImage.Set(colormapper->GetOutput());
}
catch (const itk::ExceptionObject & e)
{
std::cerr << e << std::endl;
return EXIT_FAILURE;
}
*/
return EXIT_SUCCESS;
}
`
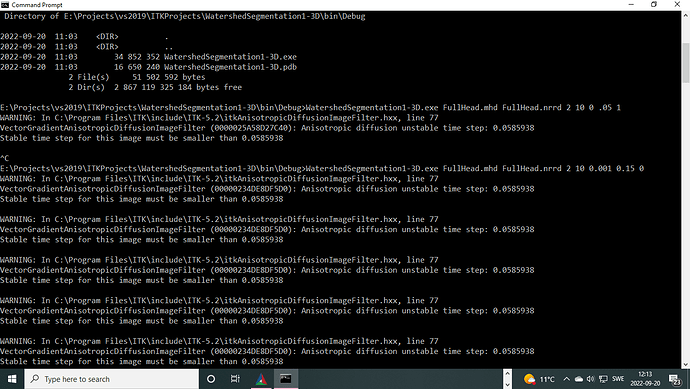
here is the screen shot of the crash for the code listing above with all the comments in that.
I already appreciate your help.
BR
@sag