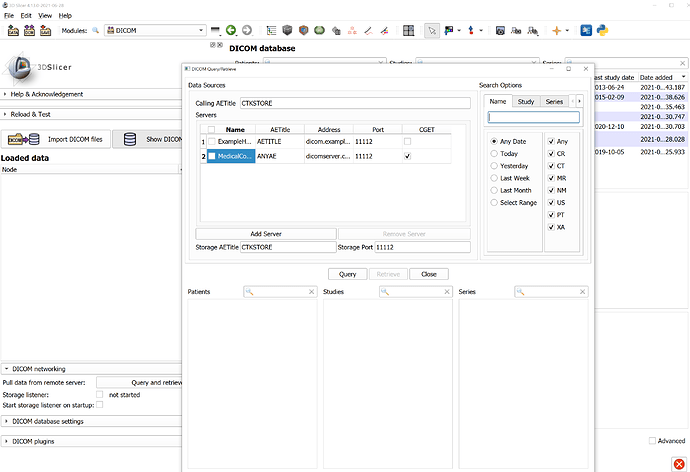
DICOM’s are uploaded in orthanc server. I need to take the DICOM’s from the server as a source object in itk.
ITK doesn’t support the communications portion of DICOM, only the file format. So you would have to download the DICOM images to files and then you could read the file using ITK.
You should be able to access your PACS using Slicer:
Alternatively, you could try to set up parameters for storescu.exe yourself.
Once you load them into Slicer, you could save them in a more convenient format, such as NRRD.
In 3D Slicer you can use classic DICOM DIMSE networking query/retrieve as @dzenanz showed in the screenshot above, but recently we also added a DICOMweb browser, which allows query/retrieve and local filtering and caching. See some more information on the DICOMweb features here. Data upload is supported, too, both via DICOMweb and DIMSE (C-store) protocol, for not just images but all information objects (segmentation, registration, radiation therapy plan, dose, etc).
Thanks for the responses.
Orthanc server (DICOM server) has restful api, so I wanted to use that
By using URL I wanted to get DICOM’s directly to the program.
As the other pointed out, there is no direct way to read DICOM directly from PACS (tcp sockets) let alone Orthanc REST api (http/https). You do need to fetch to local temporary storage and then import using the built-in GDCM library. To perform query-retrieve you do need some TCP/HTTP library in C++ ( like boost asio in combination with beast) where you can find an example for the rest api here: REST API of Orthanc — Orthanc Book documentation
Only then you can fetch locally using the following example: REST API of Orthanc — Orthanc Book documentation
Keep in mind there are multiple ways you can do that. This is simply the most straightforward workflow given by the example.
Long story short, there is nothing that easy nor straightforward when it comes to PACS and DICOM. While ITK has good DICOM support ( the file format) thanks to the built in GDCM lib, PACS is a network specification and ITK does not have ready available infrastructure for it. T
hus you need to bake it yourself with the help of 3rd party net libs. Also you need to understand DICOM and the need to filter valid image files and sort in order of Z slices, you use GDCM for that.
Orthanc’s REST API was probably implemented in the pre-DICOMweb world. I would not recommend locking yourself into it.
Instead, you can use DICOMweb, which is a REST API, too, and it is not specific to Orthanc. If you want, you can swap the storage backend to a much simpler server (like this) for a local install, or download/upload data to various web-based systems, such as Kheops (the “dropbox for DICOM images”) and Google/Azure/AWS healthcare cloud, or connect to any PACS systems (I think they all support DICOMweb now).
The DICOMweb-client Python package makes it really easy to query and retrieve or upload DICOM data sets.
OrthanC is DICOMweb compatible via a plugin: Orthanc - DICOM Server
Albite coming with a more restrictive license and not sure how complete it is either. But let me argue that is less of importance. Most hospitals (2021) will still use some sort or another of proprietary native PACS, which is not DICOMweb compliant. Therefore OrthanC REST API offers a modern way to communicate with native PACS.