HI,
I have a 3D image stacked with 2D images. I want to crop the image in the middle. Can you give me some examples to do it using ITK?
HI,
I have a 3D image stacked with 2D images. I want to crop the image in the middle. Can you give me some examples to do it using ITK?
Cropping is similar in 2D and 3D, so this 2D example should be helpful.
I wish there were better documentation for SimpleITK. I am cropping a volume after resampling and I have the bounding box dimensions that I want. However CropImageFilter requires the both the lower and upper boundaries to be the sizes of the regions rather than the bounding box.
For example,
If I have an image of size (340,340, 20) and I want to extract the region delimited by the bounding box {[10,10,0],[320,320,20]}
Then, I need to pass along the correct upper bound by doing:
[340, 340, 20] - [320, 320, 20] = [20, 20, 0] so:
crop = sitk.CropImageFilter()
crop.SetLowerBoundaryCropSize([10,10,0])
crop.SetUpperBoundaryCropSize([20,20,0])That would give me the correct area. I know it the method says it clearly “Crop Size” but to be honest it would be a lot easier to have one method where you can pass the bounding box.
Really, it was confusing.
Not sure why you are using the crop filter in SimpleITK vs. the slicing operator which I believe is what you want to do:
img[10:319, 10:319, 0:19]
With respect to learning how to work with SimpleITK we maintain an extensive set of Jupyter notebooks which we recommend people skim through before starting to use the toolkit.
Our latest tutorial (ISBI’18) is also available online, and is possibly the way to go for a quick introduction to SimpleITK.
This solution is for regular ITK.
I have been using this set of scripts so many times for cropping, and montage back 3D images, you can find them here. But the link might not be reliable in the long term, so I am copying the relevant parts:
extract_image.py
import itk
import sys
import os
if len(sys.argv) != 9:
print("Usage: " + sys.argv[0] + "inputImage outputImage index.x index.y index.z size.x size.y size.z")
sys.exit(1)
print("extract_image_filter %s" % sys.argv[1])
print("output: %s" % sys.argv[2])
input_filename = sys.argv[1]
basename = os.path.basename(input_filename)
filename = os.path.splitext(basename)[0]
output_filename = sys.argv[2]
index_x = sys.argv[3]
index_y = sys.argv[4]
index_z = sys.argv[5]
size_x = sys.argv[6]
size_y = sys.argv[7]
size_z = sys.argv[8]
reader = itk.ImageFileReader.New(FileName=input_filename)
reader.Update()
image = reader.GetOutput()
cropper = itk.ExtractImageFilter.New(Input=image)
cropper.SetDirectionCollapseToIdentity()
extraction_region = cropper.GetExtractionRegion()
size = extraction_region.GetSize()
size[0] = int(size_x)
size[1] = int(size_y)
size[2] = int(size_z)
index = extraction_region.GetIndex()
index[0] = int(index_x)
index[1] = int(index_y)
index[2] = int(index_z)
extraction_region.SetSize(size)
extraction_region.SetIndex(index)
cropper.SetExtractionRegion(extraction_region)
itk.ImageFileWriter.New(Input=cropper, FileName=output_filename).Update()
If you want to split a 3D image of size 100 100 20 by half, in for example the z direction:
python extract_image.py input_image output_image 0 0 0 100 100 10
python extract_image.py input_image output_image 0 0 10 100 100 10
Or in the second case, you pass the origin index, and the size:
python extract_image.py input_image output_image 10 10 0 310 310 20
For batch extraction, this is, splitting a 3D image in regions with no overlap. I use it because sometimes I don’t have enough RAM to compute it at once, or to speed up linear algorithms.
#!/bin/bash
# Usage batch_extract_image_filter input sizex sizey sizez geomx geomy geomz
# Example:
# let input size be : 20,20,20
# batch_extract_image_filter input 10 10 10 2 2 2
# will divide input image (20x20x20) into eight (2x2x2) images of 10x10x10 with no overlap
# There is no checking of valid sizes.
# function itk_crop
# {
input=${1}
base_name="${input%.*}"
suffix="${input##*.}"
sizex=${2}
sizey=${3}
sizez=${4}
geomx=${5}
geomy=${6}
geomz=${7}
indx=0
indy=0
indz=0
printf "input: $input \nsuffix: $suffix \n"
printf "sizes: $sizex , $sizey , $sizez\n"
printf "geoms: $geomx , $geomy , $geomz\n"
linear_index=0
z=0
while [ $z -lt $geomz ]; do
y=0
while [ $y -lt $geomy ]; do
x=0
while [ $x -lt $geomx ]; do
linear_index=$(($linear_index+1))
# echo $linear_index
echo python extract_image_filter.py $input ${base_name}_${geomx}x${geomy}x${geomz}_tile_${linear_index}.${suffix} \
$((${indx}+${sizex}*${x})) \
$((${indy}+${sizey}*${y})) \
$((${indz}+${sizez}*${z})) \
$sizex \
$sizey \
$sizez
python extract_image_filter.py $input ${base_name}_${geomx}x${geomy}x${geomz}_tile_${linear_index}.${suffix} \
$((${indx}+${sizex}*${x})) \
$((${indy}+${sizey}*${y})) \
$((${indz}+${sizez}*${z})) \
$sizex \
$sizey \
$sizez
x=$(($x+1))
done
y=$(($y+1))
done
z=$(($z+1))
done
And finally, to montage back the image: montage_tiles.py
import itk
import sys
import os
if len(sys.argv) < 4:
print("Usage: " + sys.argv[0] + "outputImage geom.x geom.y geom.z inputImage1 inputImage2 ...")
sys.exit(1)
print("montage_tiles %s" % sys.argv[1])
output_filename = sys.argv[1]
geom_x = sys.argv[2]
geom_y = sys.argv[3]
geom_z = sys.argv[4]
required_images=int(geom_x) * int(geom_y) * int(geom_z)
input_filename = sys.argv[5]
reader = itk.ImageFileReader.New(FileName=input_filename)
reader.Update()
image = reader.GetOutput()
montager = itk.TileImageFilter.New(Input=image)
layout = montager.GetLayout()
layout[0] = int(geom_x)
layout[1] = int(geom_y)
layout[2] = int(geom_z)
montager.SetLayout(layout)
# More images:
print(required_images)
for im in range(1,required_images,1):
input_filename = sys.argv[im + 5]
reader = itk.ImageFileReader.New(FileName=input_filename)
reader.Update()
montager.SetInput(im, reader.GetOutput())
itk.ImageFileWriter.New(Input=montager, FileName=output_filename).Update()
You will use it as:
python montage_tiles.py output.nrrd 2 2 2 $(for t in {1..8}; do echo /path/tiled_image_2x2x2_tile_${t}.nrrd; done)
Hope you find some bits useful!
The RegionOfInterestImageFilter may have the paramterization of the operation you desire.
Difficult to tell from the documentation. I only see SetIndex and SetSize as possible methods to use but the documentation is really bad.
Diego
Yes a bounding box or region of interest can be defined by an index and a size.
How are you wanting to define the region you are extracting?
Why is the documentation “really bad”? The documentation is take from ITK. What are you expectations? Your comment does not help the community know what is needed.
Thanks
Brad
Hi Brad,
Have you taken a look at the documentation of the class you referred to?
The documentation is really bad!! Check the description of the index method:
“odo the internal setting of the method need work!!” -> bad documentation
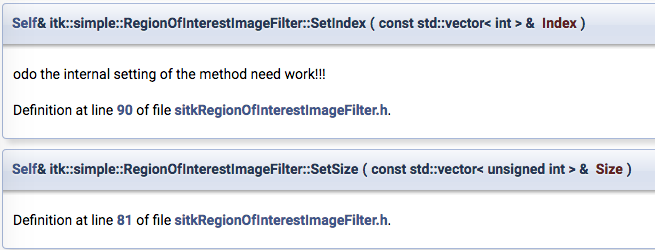
Do you think that from these docs you can honestly understand how to use the index and the size methods?
I do believe that if you point out something that is not clear or badly documented that helps the community!
Regards,
Diego
Thank you for pointing out that one item. I saw others but I did not know what your problem is.
Vague criticism is not helpful. People have worked very hard on projects and are limited by time and resources. There are many things to work on to improve a toolkit. Just saying something is “very bad” does not give any information on how to improve i lt. It is just a negative comment. I can not create an issue based on your initial feedback that could be worked on.
If you have specific feedback I please create a issue in the SimpleITK github issue tracker: Issues · SimpleITK/SimpleITK · GitHub
Thank you for your engagement and input to the ITK community.
I understand and I did not mean it as a vague or as an unappreciative criticism. I am just stating a fact. I hope that my last message provided some more clarity about this particular issue.
If I knew more about ITK I would volunteer to improve its documentation. The doxygen comments are inexistent in some cases and some classes do not have examples (especially python examples). Don’t get me wrong, I am aware this is a mammoth size effort and that there is a lack of resources. However if there was a concerted effort of improving the documentation/examples (Maybe a document-hackaton) I believe ITK would be reachable to more people and you would see more code contributions which would benefit the community!
We need to recognize the shortcomings to address them and make the software better.
Thanks,
Diego