I’m using ITK for spine reconstruction. I have some spine segmentation images in NIFTI format, how can I use ITK to reconstruct 3D spine mesh from these segmentation nifti images?
See this example and BinaryMask3DMeshSource’s documentation.
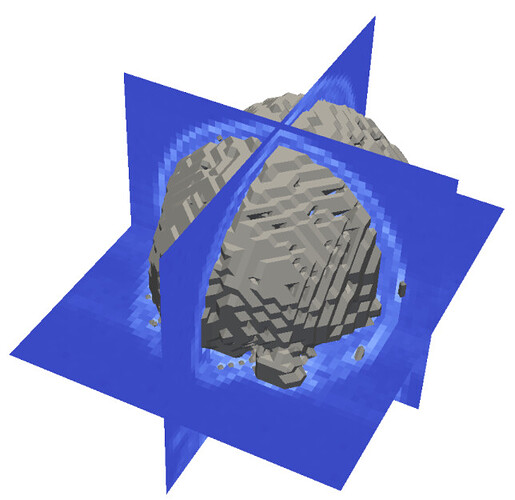
In the example I was surprised to see this mesh:
This does not look pretty. I see that the voxels are very large, but regardless of that, the constructed surface must be smooth. It seems that a very important step is missing from this example: reconstruction of the continuous signal from the discrete samples. It can be either done by processing the image before the meshing (for example, using simple low-pass filtering or some level-set based reconstruction); or can be done by smoothing the resulting mesh.
@dzenanz What ITK image or mesh filters would you recommend to reconstruct a better quality surface?
The usual choice is ITKCuberille remote module. It needs a padding filter if the object touches the edges, or else it crashes.
We recently wanted to generate quadrilateral mesh to use it in finite-element modeling as a shell (because they are more deformable than triangular meshes). It would be interesting to try if the ITKCuberille module can be used for this, but I could not find any Python examples (not even C++ examples, but I would prefer Python for quick testing anyway). I’ve checked:
Here is a Python example:
It is currently in a WIP branch in my fork, but it should eventually make it into GitHub - KitwareMedical/HASI: High-throughput Applications for Skeletal Imaging.