Hello,
I am using readimageDICOMFileSeries with the code as below. Everything worked perfectly when I had dicoms with data in UINT8 arrays. When I tested it with images with intensity values more than 255, I also received object with data encoded with 8 bytes, i.e.
{
“dimension”: 3,
“componentType”: “uint8_t”,
“pixelType”: 1,
“components”: 1
}
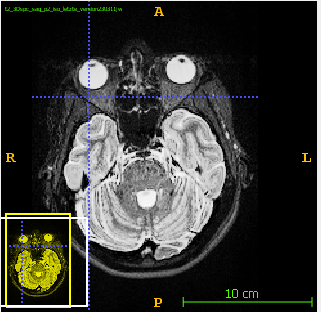
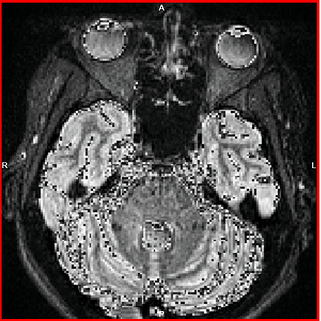
and all the points with intensity over 255 (as i checked in ITK-snap) were having wrong values (see pictures below).
My code for reading series :
function(extractedFiles) {
files= extractedFiles.map((el)=>{return new File([el.buffer],el.name)});
itkreadImageDICOMFileSeries(null, files)
.then(({ image: itkImage, webWorker }) => {
console.log(itkImage.imageType) // result as above
const vtkImage = vtkITKHelper.convertItkToVtkImage(itkImage);
//...ctd
})
.catch((err)=>{
console.log(err);
dispatch(errorFunc("Can't read ITK",imageId))});
},
}
How can I handle different encoding?
Thank you for help
Andrzej