Hello,
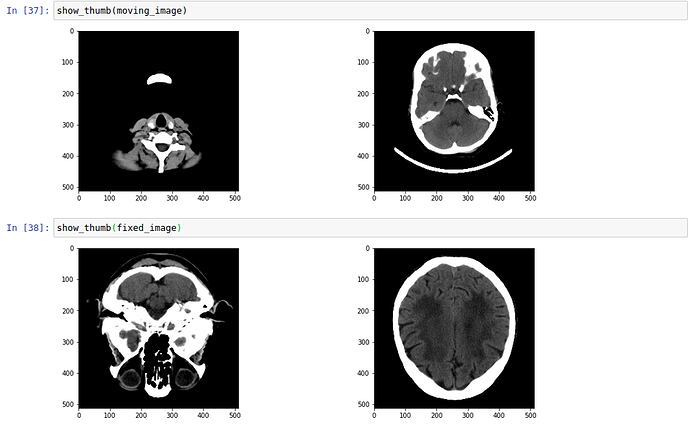
I am trying to register a head CT (with headset) on a head CT template. The presence of a headset is challenging the whole registration step (I think). It can eventually succeed if I wait long enough.
I am looking for advice on how to make the process more robust and faster, based on my current configuration.
Fixed image config :
Size: (512, 512, 55)
Spacing: (0.3547742962837219, 0.3547742962837219, 2.438863754272461)
Origin: (92.43216705322266, 83.35346984863281, -12.921518325805664)
Direction: (-1.0, 0.0, 0.0, 0.0, -1.0, 0.0, 0.0, 0.0, 1.0)
Moving image config :
Size: (512, 512, 55)
Spacing: (0.474609375, 0.474609375, 5.0)
Origin: (-121.5, 14.5, -4.4000244140625)
Direction: (1.0, 0.0, 0.0, 0.0, 1.0, 0.0, 0.0, 0.0, 1.0)
initial_transform = sitk.CenteredTransformInitializer(fixed_image,
moving_image,
sitk.Euler3DTransform(),
sitk.CenteredTransformInitializerFilter.GEOMETRY)
# registration
registration_method = sitk.ImageRegistrationMethod()
registration_method.SetMetricAsMattesMutualInformation(numberOfHistogramBins=50)
registration_method.SetMetricSamplingStrategy(registration_method.RANDOM)
registration_method.SetMetricSamplingPercentage(0.1)
registration_method.SetInterpolator(sitk.sitkLinear)
registration_method.SetOptimizerAsGradientDescent(learningRate=1.0, numberOfIterations=200)
registration_method.SetOptimizerScalesFromPhysicalShift()
# Set the initial moving and optimized transforms.
optimized_transform = sitk.Euler3DTransform()
initial_transform = sitk.CenteredTransformInitializer(sitk.Cast(fixed_image,moving_image.GetPixelID()),
moving_image,
sitk.Euler3DTransform(),
sitk.CenteredTransformInitializerFilter.GEOMETRY)
registration_method.SetMovingInitialTransform(initial_transform)
registration_method.SetInitialTransform(optimized_transform)
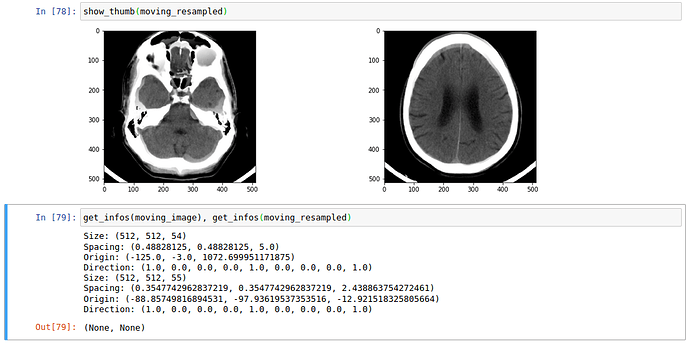
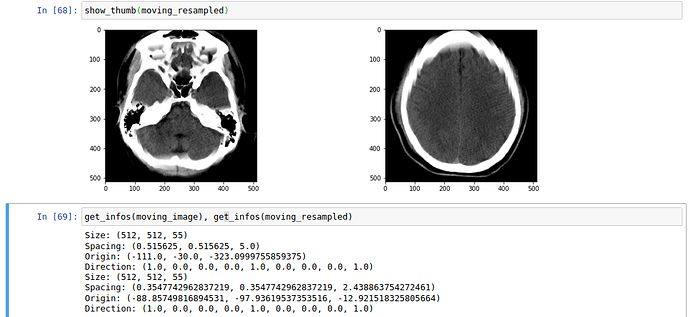
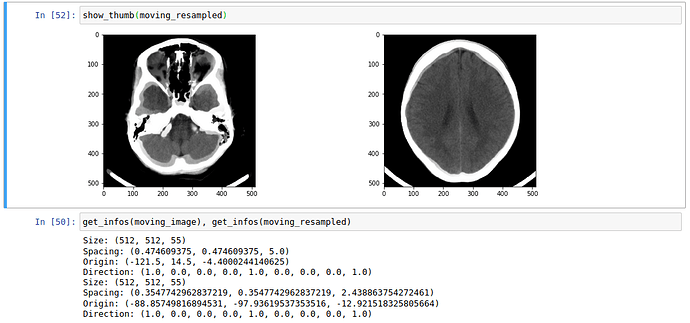
The optimization process looks good to me, though in my opinion it seems pretty long regarding the simple task I want to solve.
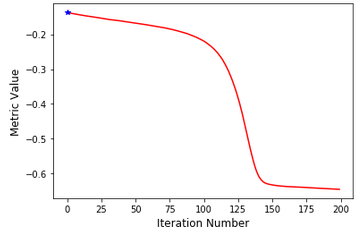
I noticed that the fixed image is not in the same orientation as the moving image, I guess that having the same orientation will help a bit. I am not sure if the complexity to register comes from the parameters or the difference in the images.
I would really appreciate some guidance 
Thanks